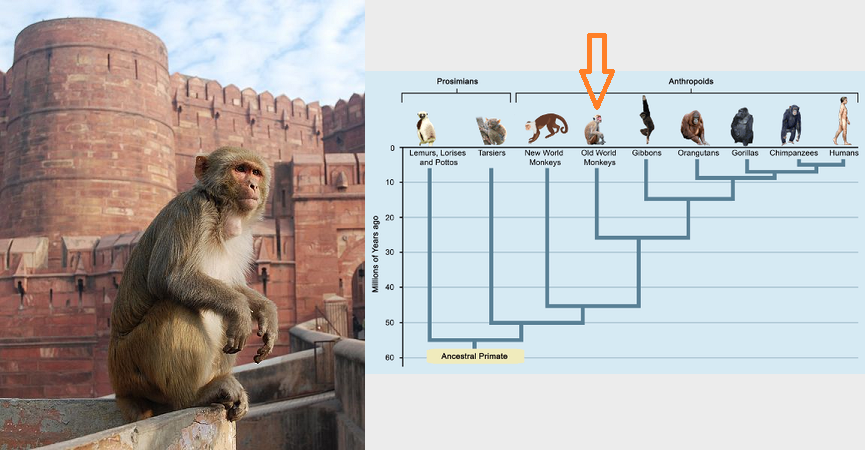
Today, we're going to use what we've learned about the command line to perform some basic bioinformatics tasks on a sample of 32 rhesus macaque whole genomes. Rhesus macaques are an Old World monkey, found throughout southern Asia, and are a common model organism for the study of human disease and primate evolution. We sequenced these genomes to study the evolution structural variation over different timescales (Thomas et al. 2020).
As a very brief overview, we sequenced genomes from 32 macaque individuals from several pedigrees and mapped the reads to the macaque reference genome. Then, using programs that look at the orientation of the mapped reads as well as the read depth, we identified regions in each sample with may have been deleted or duplicated. Today, we'll look at these structural variant calls.
You'll find in your project repo (folder) a file at the following path: data/macaque-svs/macaque-svs-filtered.bed
Note that this file has the extension .bed. This tells us something about how the text within the file is formatted.
If you remember our central dogma for Unix commands (formatted text -> command -> processed text), you'll know that being familiar
with how our files are formatted is really important.
So what is a .bed file? Bed files are used to indicate regions within a genome. It
is an extremely flexible format -- these regions can represent anything.
In it's most basic form, a .bed file consists of three columns of text, separated by a tab character. The first
column represents the chromosome or assembly scaffold of the region, while the second indicates the starting coordinate, and the third indicates
the ending coordinate of the region.
Each row represents a separate region
Additional columns can be defined, particularly of interest is a fourth column, which indicates a name or ID for the current region.
See these links for more information about .bed files:
Let's take a look at our .bed file:
less -S data/macaque-svs/macaque-svs-filtered.bed| Command line parameter | Description |
|---|---|
| less | A Linux text file viewer (use space to scroll down, b to scroll up, and q to exit) |
| -S | Turn off line-wrapping within less (use the left and right arrow keys to scroll left and right). |
| data/macaque-svs/macaque-svs-filtered.bed | The path to the file you want to view |
You should see something like this:
chr1 89943 90471 chr1:89943:<DUP>:528:1907.19
chr1 130740 131675 chr1:130740:<DEL>:935:285.63
chr1 218574 219534 chr1:218574:<DUP>:960:5699.01
chr1 219608 220078 chr1:219608:<DUP>:470:2074.69
chr1 519434 541582 chr1:519434:<DUP>:22148:1673.64
chr1 519473 542033 chr1:519473:<DUP>:22560:2560.16
chr1 520173 541800 chr1:520173:<DEL>:21627:2955.11
chr1 525401 525806 chr1:525401:<DEL>:405:2986.21
chr1 541132 590572 chr1:541132:<DEL>:49440:316.41
chr1 552968 582234 chr1:552968:<DUP>:29266:189.32
chr1 766381 766933 chr1:766381:<DEL>:552:5099.0
As defined above, this bed file has text in columns, separated by tabs. The first column being the chromosome of the region, second
being the start coordinate, and third being the end coordinate. This .bed file also has the optional
fourth column giving us a name or ID for each region. In this case, the name indicates whether the structural variant call is a
deletion (<DEL>) or a duplication (<DUP>).
So our collaborator hands us this BED file, and we want to get a general idea about the called variants. What can we do from the command line?
TASK 1: How many structural variants are there?
The most basic thing we'll want to know is how many structural variants have been called. Recalling that each line in a
bed file represents one region, which in this case means one structural variant, we can simply count the number of lines
in the file with the wc command:
wc -l data/macaque-svs/macaque-svs-filtered.bed| Command line parameter | Description |
|---|---|
| wc | The Linux word count command. Counts the number of lines, words, and characters in an input file. |
| -l | Tells wc to only count number of lines |
| data/macaque-svs/macaque-svs-filtered.bed | The path to the file you want to count |
When I do this, I see
3646Which means there are a total of 3,646 structural variants called in our sample of 32 macaques.
TASK 2: How many DELETIONS are there?
In order to answer this question, we must first separate out the deletions from the duplications. Then we can take the resulting text
and use wc -l again.
As with many problems in data science and scripting, there are many possible solutions. What we'll use is the Unix pattern matching
command, grep along with the piping function we learned about before.
grep stands for "globally search for a regular expression and print matching lines," which is a pretty descriptive way of saying it is a command for string and pattern matching. You supply grep with a file you want to search, and a string within that file you want to search for, and it will print out each line in the file that contains that string:
grep "<string or pattern>" <input file>| Command line parameter | Description |
|---|---|
| grep | A Unix string and pattern searching command |
| '<string or pattern>' | The string or pattern you want to search for |
| <input file> | The path to the file you want to search |
Searching an input file may seem like a simple task, but can be enormously helpful, and is made even more powerful by the use
of regular expressions. Regular expressions
are a syntax for string matching using wildcards to indicate patterns. The * wild card is a regular
expression that you've learned about previously which means "match ANY string."
Using regular expressions, you can both expand and narrow the types of strings you're searching for. For instance, let's say I have
a file, sample-ids.txt that lists sample IDs, one per line, all of the pattern. Some samples are
from batch "A" and follow the pattern A-
grep "A-*" sample-ids.txt| Command line parameter | Description |
|---|---|
| grep | A Unix string and pattern searching command |
| A-* | The string or pattern you want to search for. In this case, grep
will print any lines that contain the string "A-" followed by any other characters using the *
wildcard. |
| sample-ids.txt | The path to the file you want to search |
Regular expressions can get much more complex to search for more complex patterns, but for now, let's focus on the basics of
grep and how it can help us answer how many deletions are in our bed file.
As mentioned above, will use a combination of grep, piping with |, and
our line counting command, wc. Let's first just run the pattern matching command to print out
lines in our input file that represent deletions:
grep "<DEL>" data/macaque-svs/macaque-svs-filtered.bed| Command line parameter | Description |
|---|---|
| grep | A Linux string search and pattern matching command that takes as input a file or stream from a pipe and searches for a given string. |
| "<DEL>" | The pattern or specific string we want to search for |
| data/macaque-svs/macaque-svs-filtered.bed | The path to the file you want to search |
This should print out a whole lot of lines from our input file to the screen, as is the default behavior for most Unix commands. On my screen, the end of this output looks like this:
chrX 135682628 135718741 chrX:135682628:<DEL>:36113:4892.62
chrX 137821409 137821980 chrX:137821409:<DEL>:571:17208.79
chrX 146387029 146422365 chrX:146387029:<DEL>:35336:4990.42
chrY 4875713 4904730 chrY:4875713:<DEL>:29017:9217.23
chrY 4902981 4904657 chrY:4902981:<DEL>:1676:1876.99
chrY 6907076 6907624 chrY:6907076:<DEL>:548:578.79
chrY 7143528 7144240 chrY:7143528:<DEL>:712:915.06
chrY 7424851 7429392 chrY:7424851:<DEL>:4541:1978.63
chrY 10903282 10911159 chrY:10903282:<DEL>:7877:2856.38
This is similar to what we saw when we just looked at the raw file with less, but if you scrolled
through this you'll notice that all the lines printed are deletions, with the <DEL> string. This is because grep
took out search string and only returned lines that had that string in it
Now then, how does this help us know how MANY deletions there are. Well, we have two options:
-
We could save the output from grep to a file with a redirect (
>) and then usewc -lon that file. -
We could directly pipe (
|) the output fromgreptowc -l.
Let's try option number two, with piping:
grep "<DEL>" data/macaque-svs/macaque-svs-filtered.bed | wc -l| Command line parameter | Description |
|---|---|
| grep | A Linux string search and pattern matching command that takes as input a file or stream from a pipe and searches for a given string. |
| "<DEL>" | The pattern or specific string we want to search for |
| data/macaque-svs/macaque-svs-filtered.bed | The path to the file you want to search |
| | | The pipe character means the output from the command specified before it will be used as the input tot he command specified after it |
When I do this, I see
3214This means that out of our total 3,646 structural variants, 3,214, or 88%, are deletions.
TASK 3: What is the length of each variant?
To answer this question from the command line, we'll first introduce a new command: awk
awk is slightly different from the other commands we've learned about in that it is a fully defined scripting language meant to be used in conjunction with other Unix commands. Given that, it has a much more elaborate syntax than the rest of the commands: it doesn't just have input parameters, but also accepts user defined rules and functions.
This makes awk extremely powerful, and is our first step into scripting, since we will use the awk
syntax to define our own commands.
awk can be run in two ways:
- The traditional scripting/programming way, where you write your rules in a file.
- The Unix way, where you write your rules directly in the command line. These are generally referred to as one-liners.
awk '<user-defined rules>' <input file>| Command line parameter | Description |
|---|---|
| awk | A Linux text processing language |
| '<user-defined rules>' | The user-coded program to run |
| <input file> | The path to the file on which to run the program |
Like many other Unix commands, awk processes the input text in the file line-by-line, and generally for each line (or record)
it processes one column (or field) at a time. By default, awk assumes that fields are separated
by tab characers, but this can be changed with an input option.
Fields (columns) in the file are represented by variables specified by a dollar sign ($) and then the
number of the column. For instance, the first column in a file is represented by $1, the fourth column by
$4, and so on.
As a brief example, let's say we wanted to extract only the column in our input bed file that represents the end of each region. We know
from our .bed format specifications that this is the third column. So we would type:
awk '{print $3}' data/macaque-svs/macaque-svs-filtered.bed| Command line parameter | Description |
|---|---|
| awk | A Linux text processing language |
| '{print $3}' | The user-coded program to run. In this case, we only want to print the third column of the file to the screen. |
| data/macaque-svs/macaque-svs-filtered.bed | The path to the file on which to run the program |
The last few lines of this output should be:
135715923
135718741
137821980
145552312
146422365
4904730
4904657
6907624
7144240
7429392
10911159
We can do this with any other column we want, and we can perform operations on the values in those columns as well, which
brings us to how we find out the length of each region in the .bed file.
Since awk is a fully realized scripting language, it supports basic mathematical operations. Since we have
the start and end coordinates of each region, we can just tell awk to subtract the end from the start,
giving us the length of each region:
awk '{print $3 - $2}' data/macaque-svs/macaque-svs-filtered.bed| Command line parameter | Description |
|---|---|
| awk | A Linux text processing language |
| '{print $3 - $2}' | The user-coded program to run. In this case, we are telling awk to take the value in the third column and subtract the value in the second column of each row in the file. |
| data/macaque-svs/macaque-svs-filtered.bed | The path to the file on which to run the program |
The last few lines of this output should be:
36311
36113
571
1156
35336
29017
1676
548
712
4541
7877TASK 4: What is the average length of all variants?
awk can let us do more complex operations as well (it is a full scripting language!). To calculate
an average across lines in a file, we also need to keep track of a sum over all lines in the file and make a calculation at the end:
awk '{sum += $3 - $2} END {if (NR > 0) print sum / NR }' data/macaque-svs/macaque-svs-filtered.bed| Command line parameter | Description |
|---|---|
| awk | A Linux text processing language |
| '{sum += $3 - $2} END {if (NR > 0) print sum / NR }' | The user-coded program to run. In this case, we are telling awk to keep track of the sum of all lengths and average them at the end. |
| data/macaque-svs/macaque-svs-filtered.bed | The path to the file on which to run the program |
For me this prints:
3615.02Meaning our structural variants are, on average, 3,615 bases long.
But this awk program we've written is slightly more complex. Let's break it down even more:
| Code | Purpose |
|---|---|
awk |
The main call to the awk program. This tells the terminal to use
awk to process the rest of the command.
|
' |
awk programs defined as one-liners in the command line must be surrounded
by quotes to be read as a single chunk of code.
|
{sum += $3 - $2} |
This tells awk to subtract the value in the second column of the current line
from the value in the third column and to add that difference to a variable called sum.
sum is then a cumuluative sum of the lengths of all SVs over all lines in the file.
|
END |
This indicates that this is the end of the code to be executed on a per-line basis. All following code will only be executed after all lines in the file have been processed. |
{if (NR > 0) |
A conditional. NR is an internal awk
variable that stores the number of rows/lines in the input file. So this conditional states that the code
after should only be executed if there are more than 0 rows in the input file.
This prevents any divide by zero errors in the next expression.
|
print sum / NR} |
This tells awk to take the cumulative sum of all SV lengths, saved in
the variable sum and divide it by the total number of rows in the input file,
NR and print the resulting value to the screen. Since
NR represents the number of SVs, this gives us the average length of all SVs.
|
' |
awk programs defined as one-liners in the command line must be surrounded
by quotes to be read as a single chunk of code. This is the second quote, meaning the awk
program is complete.
|
data/macaque-svs/macaque-svs-filtered.bed |
This is the name of the input file on which to run the awk program
we've written between the single quotes.
|
TASK 5: What is the average length of deletions only?
Much like we piped the output from grep into wc to count the
total number of deletions, we can pipe output from grep to awk to
perform more complex operations on a subset of input data:
grep "<DEL>" data/macaque-svs/macaque-svs-filtered.bed | awk '{sum += $3 - $2} END {if (NR > 0) print sum / NR }'| Command line parameter | Description |
|---|---|
| grep | A Linux string search and pattern matching command that takes as input a file or stream from a pipe and searches for a given string. |
| "<DEL>" | The pattern or specific string we want to search for |
| data/macaque-svs/macaque-svs-filtered.bed | The path to the file you want to search |
| | | The pipe character means the output from the command specified before it will be used as the input to the command specified after it |
| awk | A Linux text processing language |
| '{sum += $3 - $2} END {if (NR > 0) print sum / NR }' | The user-coded program to run. In this case, we are telling awk to keep track of the sum of all lengths and average them at the end. |
| data/macaque-svs/macaque-svs-filtered.bed | The path to the file on which to run the program |
So now awk, rather than having an input file specified, takes input from the output of
grep. For me, this prints the average length of deletions as:
3161.33This is pretty close to the overall average of 3,615bp, but a little shorter. What about the average length of duplications?
grep "<DUP>" data/macaque-svs/macaque-svs-filtered.bed | awk '{sum += $3 - $2} END {if (NR > 0) print sum / NR }'| Command line parameter | Description |
|---|---|
| grep | A Linux string search and pattern matching command that takes as input a file or stream from a pipe and searches for a given string. |
| "<DUP>" | The pattern or specific string we want to search for |
| data/macaque-svs/macaque-svs-filtered.bed | The path to the file you want to search |
| | | The pipe character means the output from the command specified before it will be used as the input to the command specified after it |
| awk | A Linux text processing language |
| '{sum += $3 - $2} END {if (NR > 0) print sum / NR }' | The user-coded program to run. In this case, we are telling awk to keep track of the sum of all lengths and average them at the end. |
| data/macaque-svs/macaque-svs-filtered.bed | The path to the file on which to run the program |
6990.42Wow! So deletions make up almost 90% of all SVs, but duplications are, on average, more than twice as long.
TASK 6: How many SVs overlap with genes?
For many types of genomic variation the question of how they affect annotated functional regions, like genes, will inevitably be asked. In order to answer this question, we'll need to familiarize ourselves with another file format, that of GTF and GFF files.
GTF stands for General Transfer Format and GFF stands for General Feature Format.
GTF and GFF files are both tab delimited formats that indicate regions in a
genome, much like a bed file. However, these files are limited to annotated functional regions and contain
extra information about those regions, such as functional class (e.g., gene, transcript, UTR), and orientation.
Read more about GTF and GFF file formats here:
We've pre-downloaded a GTF file for the rhesus macaque from Ensembl. Let's take a look at it:
zless -S data/macaque-svs/annotation-files/Macaca_mulatta.Mmul_8.0.1.97.chromes.gtf.gz| Command line parameter | Description |
|---|---|
| zless | A Linux compressed text file viewer (use space to scroll down, b to scroll up, and q to exit) |
| -S | Turn off line-wrapping within zless (use the left and right arrow keys to scroll left and right). |
| data/macaque-svs/annotation-files/Macaca_mulatta.Mmul_8.0.1.97.chromes.gtf.gz | The path to the file you want to view |
You should see something like this:
1 ensembl gene 25432 42232 . + . gene_id "ENSMMUG00000005947"; gene_version "3"; gene_name "SAMD11"; gene_source "ensembl"; gene_biotype "protein_coding";
1 ensembl transcript 25432 42232 . + . gene_id "ENSMMUG00000005947"; gene_version "3"; transcript_id "ENSMMUT00000063154"; transcript_version "1"; gene_name "SAMD11"; gene_source "ensembl"; gene_biotype "protein_coding"; transcript_name "SAMD11-201"; transcript_source "ensembl"; transcript_biotype "protein_coding";
1 ensembl exon 25432 25503 . + . gene_id "ENSMMUG00000005947"; gene_version "3"; transcript_id "ENSMMUT00000063154"; transcript_version "1"; exon_number "1"; gene_name "SAMD11"; gene_source "ensembl"; gene_biotype "protein_coding"; transcript_name "SAMD11-201"; transcript_source "ensembl"; transcript_biotype "protein_coding"; exon_id "ENSMMUE00000311984"; exon_version "2";
1 ensembl CDS 25432 25503 . + 0 gene_id "ENSMMUG00000005947"; gene_version "3"; transcript_id "ENSMMUT00000063154"; transcript_version "1"; exon_number "1"; gene_name "SAMD11"; gene_source "ensembl"; gene_biotype "protein_coding"; transcript_name "SAMD11-201"; transcript_source "ensembl"; transcript_biotype "protein_coding"; protein_id "ENSMMUP00000049946"; protein_version "1";
1 ensembl start_codon 25432 25434 . + 0 gene_id "ENSMMUG00000005947"; gene_version "3"; transcript_id "ENSMMUT00000063154"; transcript_version "1"; exon_number "1"; gene_name "SAMD11"; gene_source "ensembl"; gene_biotype "protein_coding"; transcript_name "SAMD11-201"; transcript_source "ensembl"; transcript_biotype "protein_coding";
1 ensembl exon 29573 29754 . + . gene_id "ENSMMUG00000005947"; gene_version "3"; transcript_id "ENSMMUT00000063154"; transcript_version "1"; exon_number "2"; gene_name "SAMD11"; gene_source "ensembl"; gene_biotype "protein_coding"; transcript_name "SAMD11-201"; transcript_source "ensembl"; transcript_biotype "protein_coding"; exon_id "ENSMMUE00000311983"; exon_version "1";
1 ensembl CDS 29573 29754 . + 0 gene_id "ENSMMUG00000005947"; gene_version "3"; transcript_id "ENSMMUT00000063154"; transcript_version "1"; exon_number "2"; gene_name "SAMD11"; gene_source "ensembl"; gene_biotype "protein_coding"; transcript_name "SAMD11-201"; transcript_source "ensembl"; transcript_biotype "protein_coding"; protein_id "ENSMMUP00000049946"; protein_version "1";
So this should have all the columns required of a GTF file as outlined in the links above. Notably, the first,
third, and fourth columns define the location within the genome, being the chromosome, start coordinate, and end coordinate respectively.
Columns 2 defines the regions annotation source while column 3 defines the type of region. You can notice here the nesting structure
associated with genes and their sub-elements.
Column 6 shows the regions orientation. This is an often overlooked but important thing to consider when extracting sequences in a genome: if the region indicated is on the opposite strand (indicated with a - sign), then you must take the reverse complement of that sequence, and for genes the sub-elements (exons, CDS, etc.) must be taken in reverse order. Today, since we're not actually extracting sequence and only looking at coordinates, we shouldn't need to worry about this.
The last column in the GTF files serves as a sort of catch-all. This column is named 'attributes', but programs
and users are free to put any information in this column that they like, separated by semi-colons. Unfortunately, this leads to variations
between GTF files, which can cause trouble for down-stream users. Again, today we don't need to worry about this.
In order to address our question, "How many SVs overalp with genes?", we'll be using a program called
bedtools. bedtools
is a staple bioinformatic program used for working with genomic regions in bed format. It is actually
a suite of tools designed with the Unix philosophy: each sub-program does one thing really well, and the sub-programs can work
together by taking input from each other.
Since bedtools works with bed files, we'll need to convert our
GTF regions of interest to bed format.
First, let's use awk to extract only lines that are defined as "gene" in the third column:
zcat data/macaque-svs/annotation-files/Macaca_mulatta.Mmul_8.0.1.97.chromes.gtf.gz | awk '{if($3 == "gene"){print}}'| Command line parameter | Description |
|---|---|
| zcat | This prints lines from a compressed text file to the screen. Since this file is compressed (.gz) we first need to decompress the text for awk to read. |
| data/macaque-svs/annotation-files/Macaca_mulatta.Mmul_8.0.1.97.chromes.gtf.gz | The path to the input file to read |
| | | The pipe character means the output from the command specified before it will be used as the input to the command specified after it |
| awk | A Linux text processing language |
| '{if($3 == "gene"){print}}' | The user-coded program to run. In this case, we want to print the line to the screen if the third column matches the string "gene". |
This should print all the lines that are genes to the screen. But now we want to convert these lines into bed
format. This should be easy since all the information we need is in the first, fourth, and fifth columns:
zcat data/macaque-svs/annotation-files/Macaca_mulatta.Mmul_8.0.1.97.chromes.gtf.gz | awk '{if($3 == "gene"){print}}' | awk 'BEGIN{OFS="\t"}{print "chr"$1, $4, $5}'| Command line parameter | Description |
|---|---|
| zcat | This prints lines from a compressed text file to the screen. Since this file is compressed (.gz) we first need to decompress the text for awk to read. |
| data/macaque-svs/annotation-files/Macaca_mulatta.Mmul_8.0.1.97.chromes.gtf.gz | The path to the input file to read |
| | | The pipe character means the output from the command specified before it will be used as the input to the command specified after it |
| awk | A Linux text processing language |
| '{if($3 == "gene"){print}}' | The user-coded program to run. In this case, we want to print the line to the screen if the third column matches the string "gene". |
| | | The pipe character means the output from the command specified before it will be used as the input to the command specified after it |
| awk | A Linux text processing language |
| 'BEGIN{OFS="\t"}{print "chr"$1, $4, $5}' | The user-coded program to run. In this case, we want to print the first, fourth, and fifth columns of the input file. |
This should print the specified columns of all lines that have "gene" in the third column to the screen. One new thing here is that
we've re-set the internal awk variable OFS (output field separator) to a tab
character. This means output columns will print with this character between them. Using a tab character matches bed
format specifications. Note that also columns must be listed with commas in the awk program.
You'll also notice that we've added a string to the first column: "chr". This is because the bed file with our SV
calls names chromosomes as chr1, chr2, etc., while this GTF file simply names them 1, 2, etc. This mis-match would
prevent bedtools from finding overlaps so we need to add the "chr" string here.
Finally, since we want to use this text later, we'll need to save it by redirecting it to a file with >:
zcat data/macaque-svs/annotation-files/Macaca_mulatta.Mmul_8.0.1.97.chromes.gtf.gz | awk '{if($3 == "gene"){print}}' | awk 'BEGIN{OFS="\t"}{print "chr"$1, $4, $5}' > data/macaque-svs/annotation-files/macaque-genes.bed| Command line parameter | Description |
|---|---|
| zcat | This prints lines from a compressed text file to the screen. Since this file is compressed (.gz) we first need to decompress the text for awk to read. |
| data/macaque-svs/annotation-files/Macaca_mulatta.Mmul_8.0.1.97.chromes.gtf.gz | The path to the input file to read |
| | | The pipe character means the output from the command specified before it will be used as the input to the command specified after it |
| awk | A Linux text processing language |
| '{if($3 == "gene"){print}}' | The user-coded program to run. In this case, we want to print the line to the screen if the third column matches the string "gene". |
| | | The pipe character means the output from the command specified before it will be used as the input to the command specified after it |
| awk | A Linux text processing language |
| 'BEGIN{OFS="\t"}{print "chr"$1, $4, $5}' | The user-coded program to run. In this case, we want to print the first, fourth, and fifth columns of the input file. |
| > | The redirect character means the output from the command specified before it will be saved to the file specified after it |
| data/macaque-svs/annotation-files/macaque-genes.bed | The path to the desired output file. This will be created if it doesn't exist and WILL OVERWRITE files that do exist |
Now, instead of printing the output of our command to the screen, it will be saved in the file
macaque-genes.bed, and we can use it as we wish later.
As we mentioned above, bedtools, is an extremely
versatile program for working with genomic regions in bed format. bedtools
is a suite of programs that are generally run as follows:
bedtools <sub-program> <options> <input file>| Command line parameter | Description |
|---|---|
| bedtools | A program for working with genomic regions in bed format |
| <sub-program> | The desired sub-program within bedtools to run |
| <options> | Sub-program specific run options |
| <input file> | The path to the file you want to run on |
A lot more info can be found on the bedtools website.
Now we're going to use bedtools intersect
to see how many of our structural variants overlap with genes. bedtools intersect takes as input 2 bed
files and prints output based on their overlaps. This output can be customized to include the number or percent of bases overlapped,
among other things.
bedtools intersect -u -a data/macaque-svs/macaque-svs-filtered.bed -b data/macaque-svs/annotation-files/macaque-genes.bed| Command line parameter | Description |
|---|---|
| bedtools | A program for working with genomic regions in bed format |
| intersect | A sub-program for bedtools that compares 2 bed files and reports overlaps |
| -u | This option says to print the region in file a if it has any overlaps in file b |
| -a | The option to specify the first bed file |
| data/macaque-svs/macaque-svs-filtered.bed | The path to the first bed file |
| -b | The option to specify the second bed file |
| data/macaque-svs/annotation-files/macaque-genes.bed | The path to the second bed file |
This prints out all lines in file a that overlap with a region in file b. The end of my output looks something like this:
chrX 86600287 86600725 chrX:86600287:<DEL>:438:6027.45
chrX 86613955 86615407 chrX:86613955:<DEL>:1452:1737.19
chrX 91174961 91176888 chrX:91174961:<DEL>:1927:4205.93
chrX 92753089 92753972 chrX:92753089:<DEL>:883:3724.23
chrX 98845605 98847132 chrX:98845605:<DEL>:1527:1678.29
chrX 123944663 123946419 chrX:123944663:<DEL>:1756:2666.16
chrX 129328746 129330969 chrX:129328746:<DEL>:2223:5690.06
chrX 135679612 135715923 chrX:135679612:<DEL>:36311:1521.72
chrX 135682628 135718741 chrX:135682628:<DEL>:36113:4892.62
chrX 146387029 146422365 chrX:146387029:<DEL>:35336:4990.42
These are the SVs in our sample that overlap genes in the macaque genome! How many of these are there? Let's pipe this output
into wc:
bedtools intersect -u -a data/macaque-svs/macaque-svs-filtered.bed -b data/macaque-svs/annotation-files/macaque-genes.bed | wc -l| Command line parameter | Description |
|---|---|
| bedtools | A program for working with genomic regions in bed format |
| intersect | A sub-program for bedtools that compares 2 bed files and reports overlaps |
| -u | This option says to print the region in file a if it has any overlaps in file b |
| -a | The option to specify the first bed file |
| data/macaque-svs/macaque-svs-filtered.bed | The path to the first bed file |
| -b | The option to specify the second bed file |
| data/macaque-svs/annotation-files/macaque-genes.bed | The path to the second bed file |
| | | The pipe character means the output from the command specified before it will be used as the input to the command specified after it |
| wc | The Linux word count command. Counts the number of lines, words, and characters in an input file. |
| -l | Tells wc to only count number of lines |
When I do this, I see
1609Meaning 1,609 structural variants overlap a gene. This is roughly 44%.
Next, we'll build on these bioinformatic skills by looking at a set of SNPs from a sample of wolves!