The PhyloAcc interface was run on Sunday Oct 23, 2022 at 10:05:47 EDT on bioinf02.rc.fas.harvard.edu as follows:
/n/home07/gthomas/anaconda3/envs/phyloacc-pkg/bin/phyloacc.py -d /n/holylfs05/LABS/informatics/Everyone/phyloacc-data/mammal-input-accelerated/seq/ -m /n/holylfs05/LABS/informatics/Everyone/phyloacc-data/mammal-input-accelerated/mammal_acc1.mod -l /n/holylfs05/LABS/informatics/Everyone/phyloacc-data/mammal-input-accelerated/tree_coal_unit.mod -t turTru2;orcOrc1;odoRosDiv1;lepWed1;triMan1 -g monDom5;sarHar1;macEug2;ornAna1 -r adaptive -burnin 1000 -mcmc 7000 -n 24 -p 8 -j 20 -batch 25 -part holy-info,shared,holy-smokes -time 24 -o /n/holylfs05/LABS/informatics/Everyone/phyloacc-data/workshop-20221027/mammal/adaptive-test-branches/ -phyloacc CONSERVE_PRIOR_B 0.1;ACCE_PRIOR_A 4; ACCE_PRIOR_B 0.5; CONSERVE_PROP 0.5; CONSERVE_RATE 0.3; TRIM_GAP_PERCENT 0.7 --overwrite
| Input alignments | 2030 |
| Alignments with 0 informative sites (removed) | 1 |
| Alignments for species tree model | 28 |
| Alignments for gene tree model | 2001 |
| Batches | 83 |
| Alignments per batch | 25 |
| Processes to use per batch | 8 |
| Batches using species tree model | 2 |
| Batches using gene tree model | 81 |
| Batches to submit to cluster simultaneously | 20 |
See the full log file for more info.
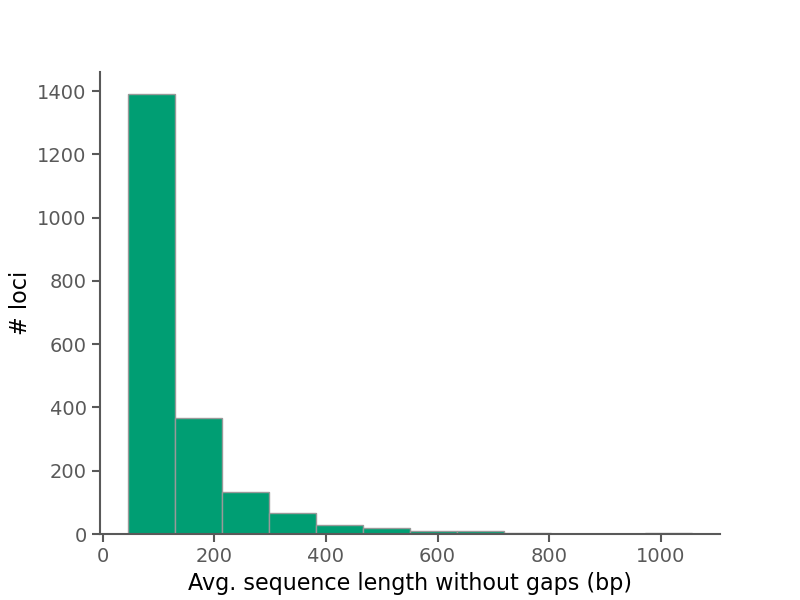
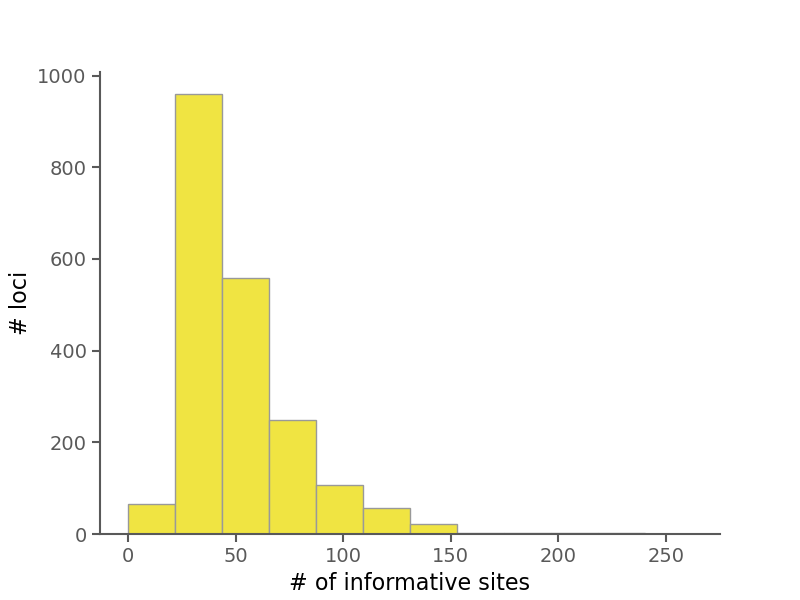
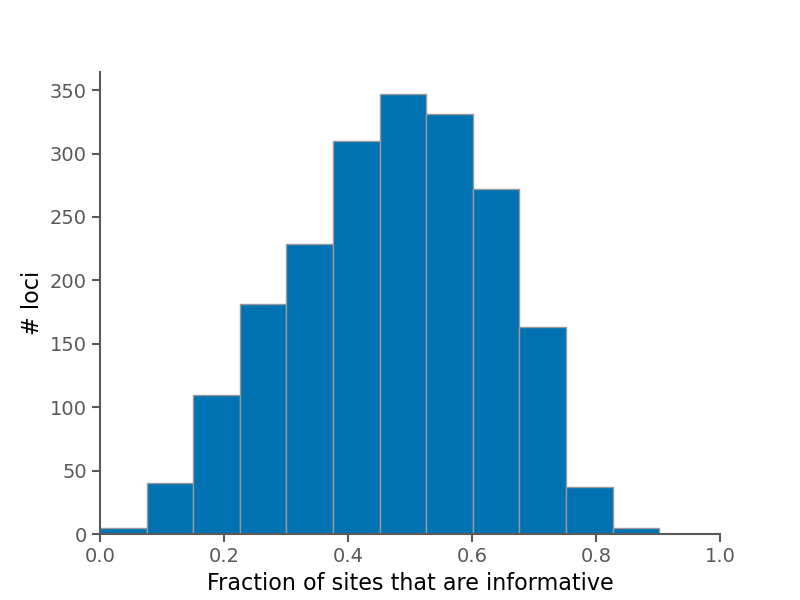
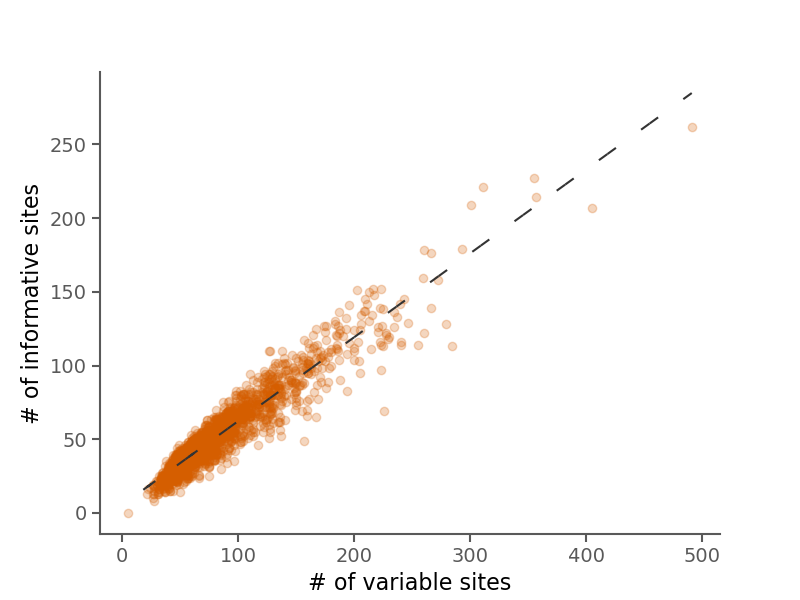
Below are some summary plots. Raw data is also available in CSV format in the alignment stats file.
The generated batches can be run with Snakemake with the following command:
snakemake -p -s /n/holylfs05/LABS/informatics/Everyone/phyloacc-data/workshop-20221027/mammal/adaptive-test-branches/phyloacc-job-files/snakemake/run_phyloacc.smk --configfile /n/holylfs05/LABS/informatics/Everyone/phyloacc-data/workshop-20221027/mammal/adaptive-test-branches/phyloacc-job-files/snakemake/phyloacc-config.yaml --profile /n/holylfs05/LABS/informatics/Everyone/phyloacc-data/workshop-20221027/mammal/adaptive-test-branches/phyloacc-job-files/snakemake/profiles/slurm_profile --dryrun
Note that this includes the --dryrun option to check for possible errors before submitting jobs. If everything looks good
after running that, feel free to remove the --dryrun option to submit the jobs to your cluster. You may also want to run
this in the background using screen,
tmux or some other terminal multiplexer
to avoid disruptions due to server disconnects.
Since some or all of the loci will be processed with the gene tree model, a species tree with branch lengths in coalescent
units was provided: /n/holylfs05/LABS/informatics/Everyone/phyloacc-data/mammal-input-accelerated/tree_coal_unit.mod
The branch lengths from this tree will be used in conjunction with the branch lengths in the input tree (which should be in units of relative number of substitutions) to estimate the theta parameter used in the gene tree model.
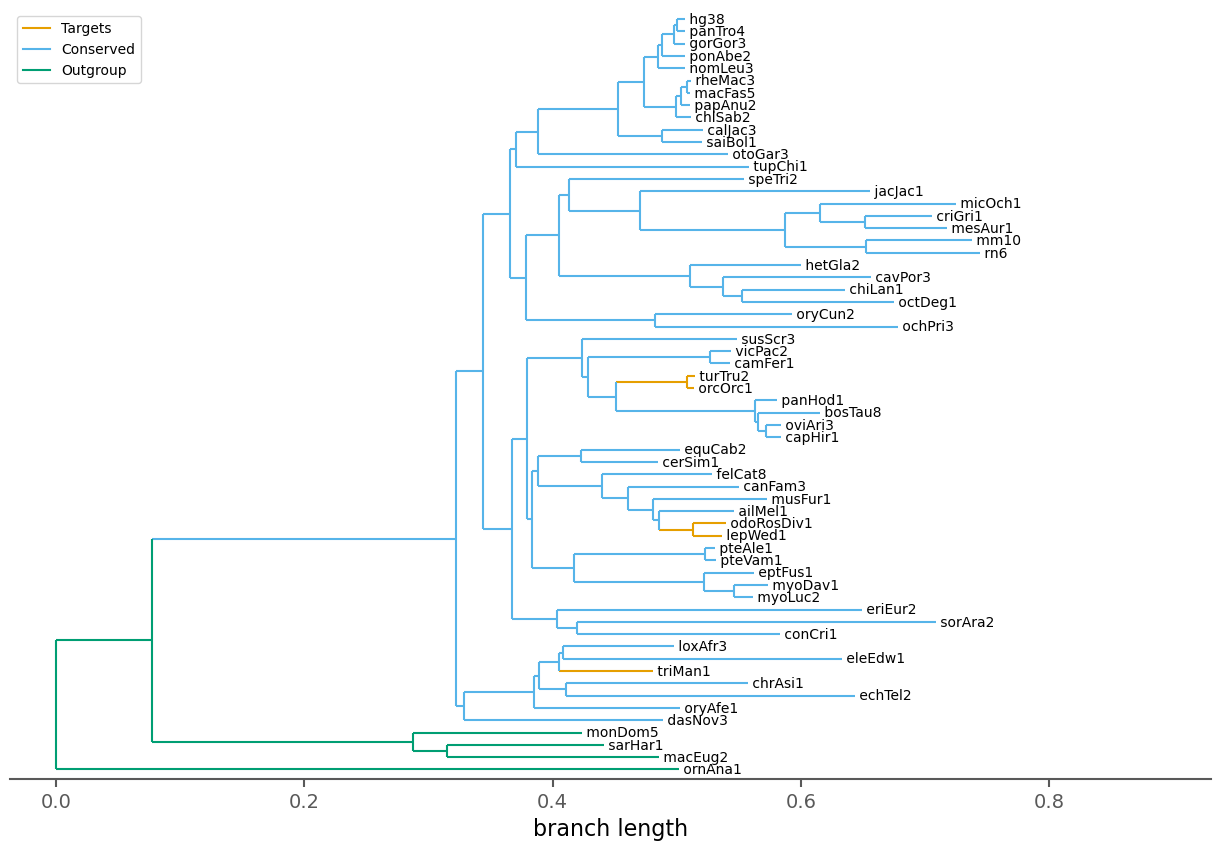
| Species | 62 |
| Target species | 5 |
| Conserved species | 53 |
| Outgroup species | 4 |
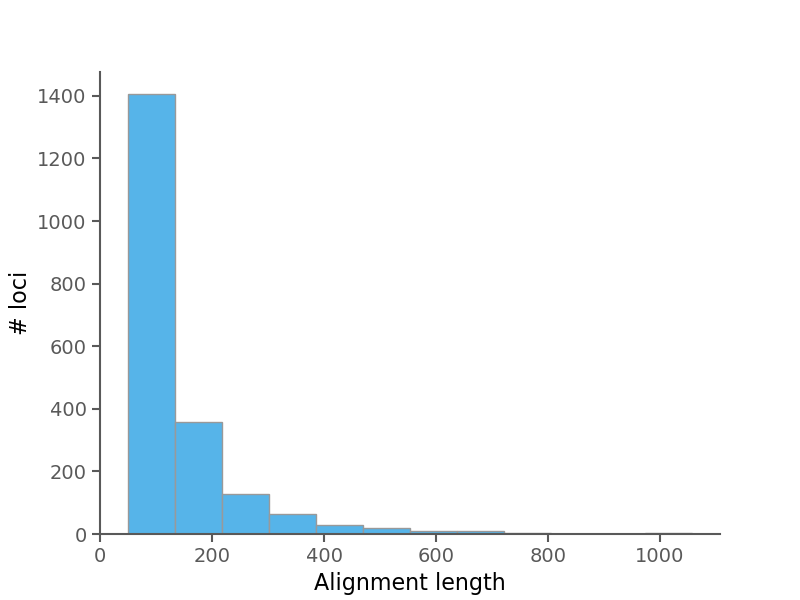
| Average alignment length | 129.627 |
| Median alignment length | 92.0 |
| Average sequence length (no gaps) per alignment | 128.71 |
| Median sequence length (no gaps) per alignment | 90.742 |
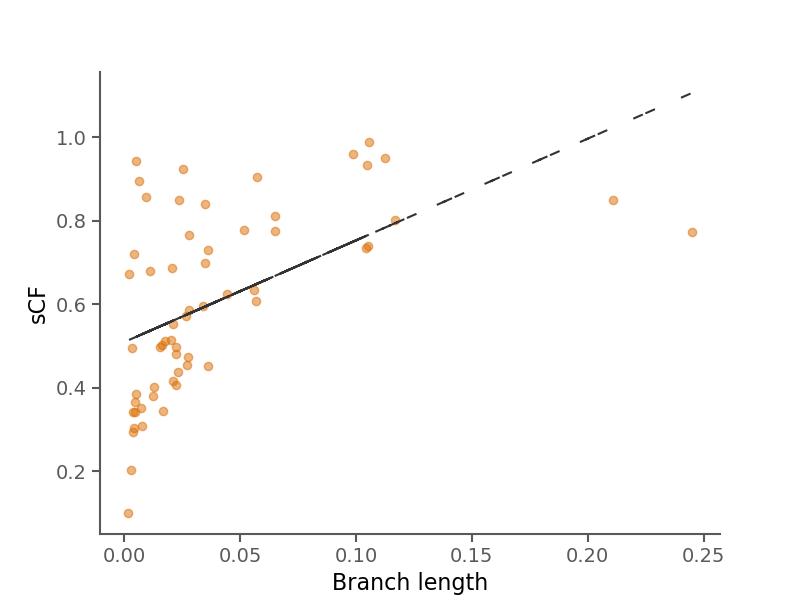
Site concordance factors (sCF) summarize the number of sites in an alignment that support a branch in a given phylogeny.
Briefly, they are computed by sampling quartets of species around each branch in an unrooted tree and counting the number of decisive sites that match the topology of the sampled quartet in the phylogeny. See Minh et al. 2020 for more information.
Here, rather than averaging over branches for all alignments, we calculate sCF for each branch in each alignment separately.
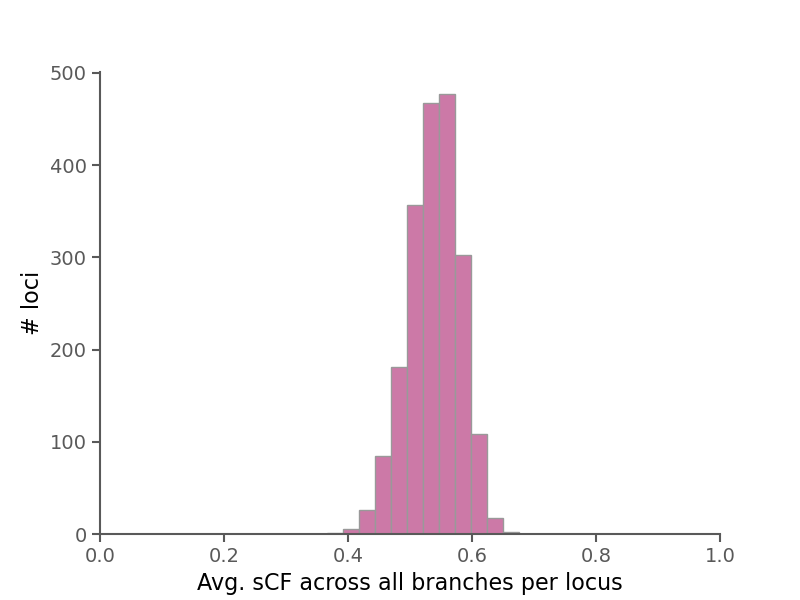
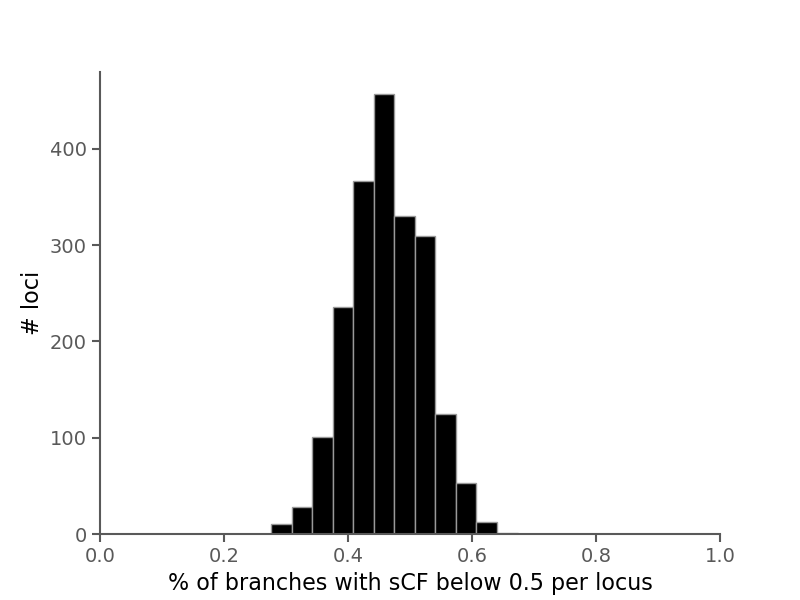
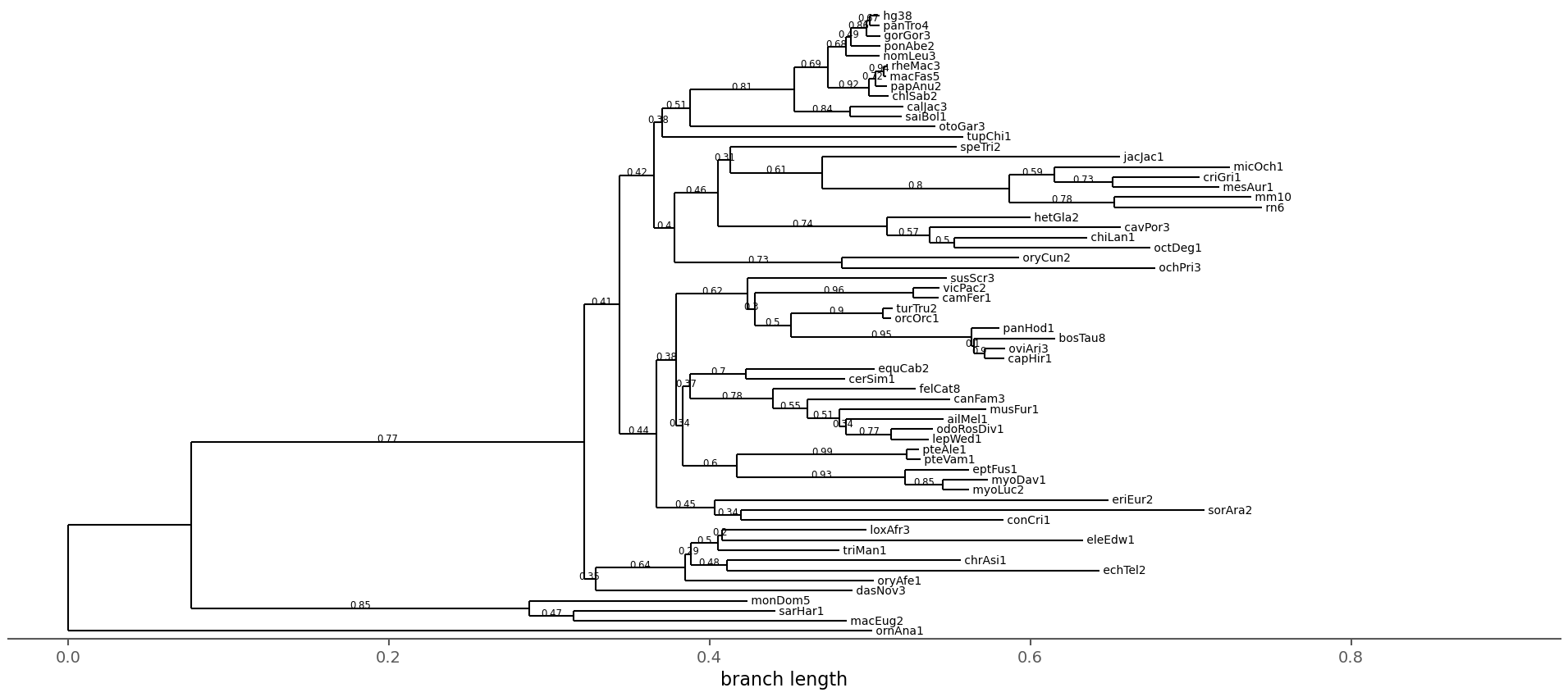
Here, the species tree is shown with branches labeled by sCF across all branches (as described in Minh et al. 2020).
The tree in Newick format is located here.
The sCF stats per branch can be found in CSV format here.